I am trying to understand the basic difference between stepwise and backward regression in R using the step function.įor stepwise regression I used the following command step(lm(mpg~wt+drat+disp+qsec,data=mtcars),direction="both") It is also useful to compile the time the document was processed. Here is the session information devtools::session_info() # setting value # 16 pdata$tissue.typewhite_blood_cell -8.40407714 NaN NaNĪnother good idea is to look for “patterns” in the residuals colramp = colorRampPalette(1:4)(17) # 14 pdata$tissue.typetestes 0.47932985 NaN NaN # 13 pdata$tissue.typeskeletal_muscle -4.27479412 NaN NaN # 12 pdata$tissue.typeprostate -0.79305234 NaN NaN # 10 pdata$tissue.typelymphnode -2.43829285 NaN NaN 
# 8 pdata$tissue.typeliver 0.52887576 NaN NaN # 7 pdata$tissue.typekidney -0.76564122 NaN NaN # 4 pdata$tissue.typebreast -0.55076758 NaN NaN # 2 pdata$tissue.typeadrenal -2.64252590 NaN NaN Tidy(lm8) # term estimate std.error statistic lm8 = lm(gene1 ~ pdata$tissue.type + pdata$age) This is also a problem if you fit too many variables for example. R fails “gracefully” (i.e. doesn’t tell you) when this happens so you have to check by hand. This is called co-linearity and can lead to highly variable and uninterpretable results. gene1 = log2(edata+1)īe careful when two variables in your regression model are very highly correlated. par(mfrow=c(1,2))ĭata transforms are often applied before regression and the residuals look a little better here. Outliers in the residuals aren’t great either.  
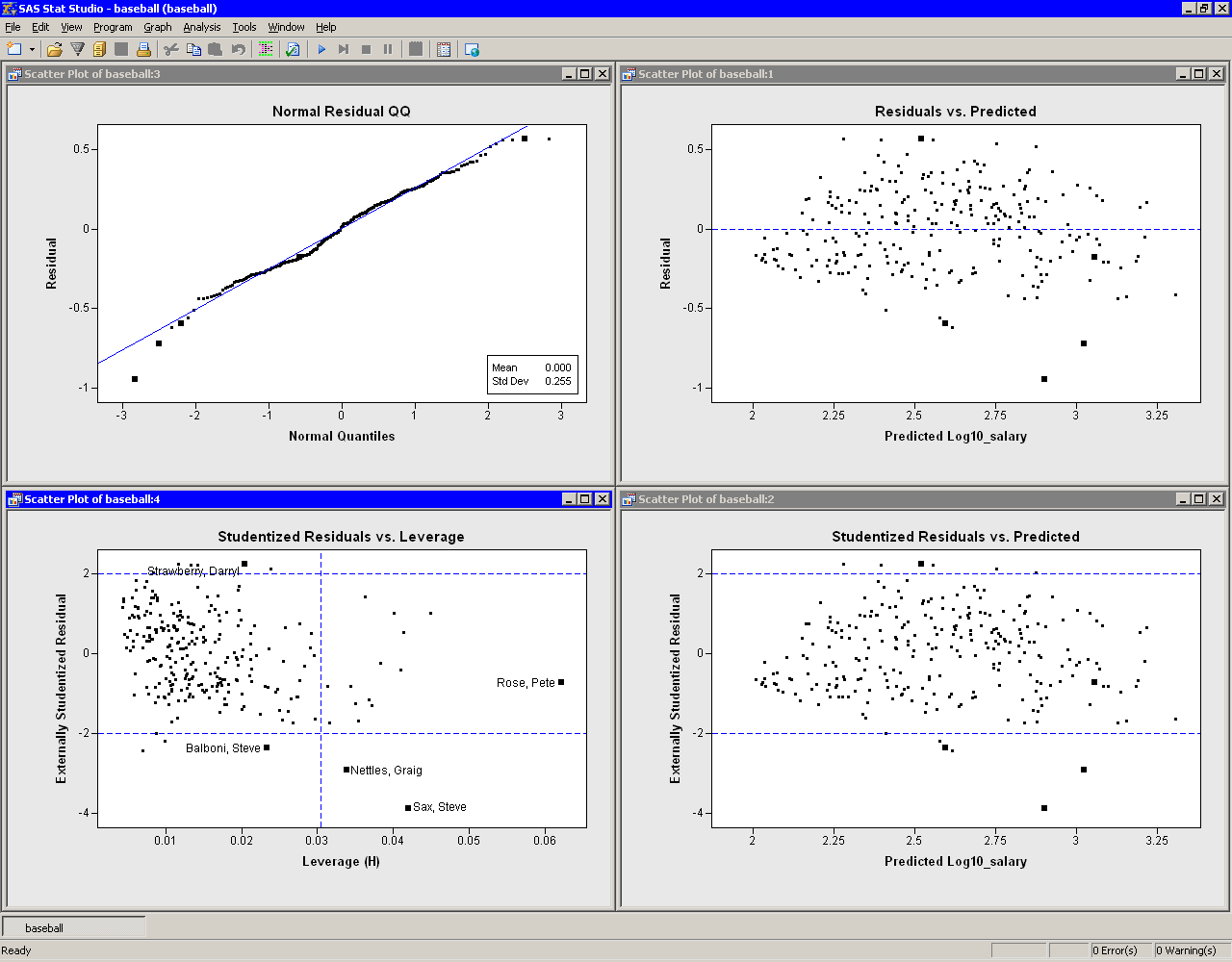
In general you’d like your residuals to looks symmetrically (e.g. approximately Normally) distributed but they aren’t here. Here there isn’t much impact lm4 = lm(edata ~ pdata$age)īut in this case there is a huge impact index = 1:19 Outliers can have a big impact on the regression, depending on where they land. This leads to multiple coefficients for one variable tidy(lm(edata ~ pdata$tissue.type )) # term estimate std.error statistic
0 Comments
Leave a Reply. |

 RSS Feed
RSS Feed
